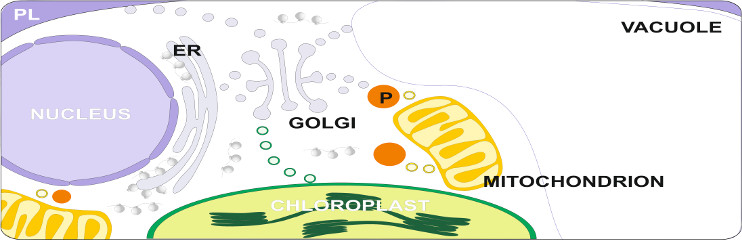
Regulation and function of heat stress transcription factors and molecular chaperones in protein homeostasis
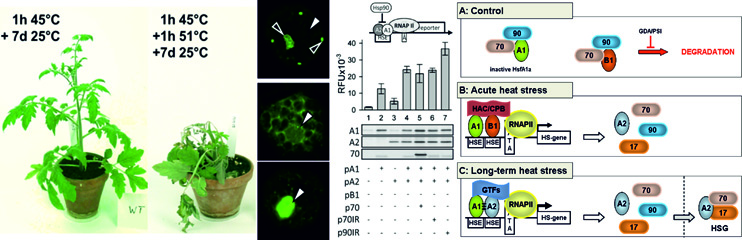
Rosenkranz RRE, Bachiri S, Vraggalas S, Keller M, Simm S, Schleiff E, Fragkostefanakis S. Front Plant Sci. 2021 Mar 29;12:645689. doi: 10.3389/fpls.2021.645689. PMID: 33854522; PMCID: PMC8039515.
Natural variation in HsfA2 pre-mRNA splicing is associated with changes in thermotolerance during tomato domestication.
Hu Y, Mesihovic A, Jiménez-Gómez JM, Röth S, Gebhardt P, Bublak D, Bovy A, Scharf KD, Schleiff E, Fragkostefanakis S. New Phytol. 2020 Feb;225(3):1297-1310. doi: 10.1111/nph.16221. Epub 2019 Nov 14. PMID: 31556121
The repressor and co-activator HsfB1 regulates the major heat stress transcription factors in tomato.
Fragkostefanakis S, Simm S, El-Shershaby A, Hu Y, Bublak D, Mesihovic A, Darm K, Mishra SK, Tschiersch B, Theres K, Scharf C, Schleiff E, Scharf KD. Plant Cell Environ. 2019 Mar;42(3):874-890. doi: 10.1111/pce.13434. Epub 2018 Oct 11. PMID: 30187931
Alternative splicing in tomato pollen in response to heat stress.
Keller M, Hu Y, Mesihovic A, Fragkostefanakis S, Schleiff E, Simm S. DNA Res. 2017 Apr 1;24(2):205-217. doi: 10.1093/dnares/dsw051. PMID: 28025318
HsfA2 Controls the Activity of Developmentally and Stress-Regulated Heat Stress Protection Mechanisms in Tomato Male Reproductive Tissues.
Fragkostefanakis S, Mesihovic A, Simm S, Paupière MJ, Hu Y, Paul P, Mishra SK, Tschiersch B, Theres K, Bovy A, Schleiff E, Scharf KD. Plant Physiol. 2016 Apr;170(4):2461-77. doi: 10.1104/pp.15.01913. Epub 2016 Feb 25. PMID: 26917685
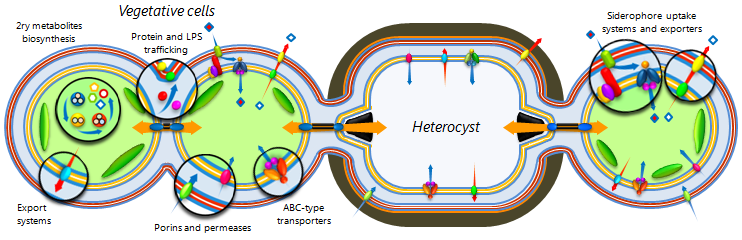
Anabaena sp. PCC7120, a model for cell differentiation and chloroplast evolution
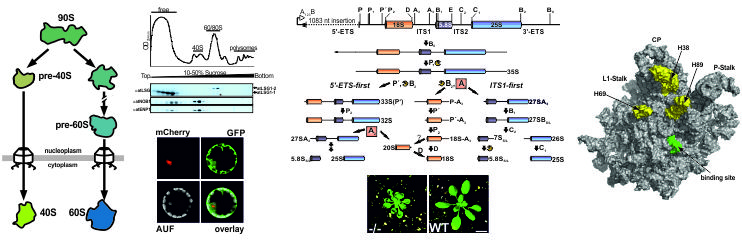
Plant Ribosome Biogenesis
Expression and structural analysis to understand evolution of plants
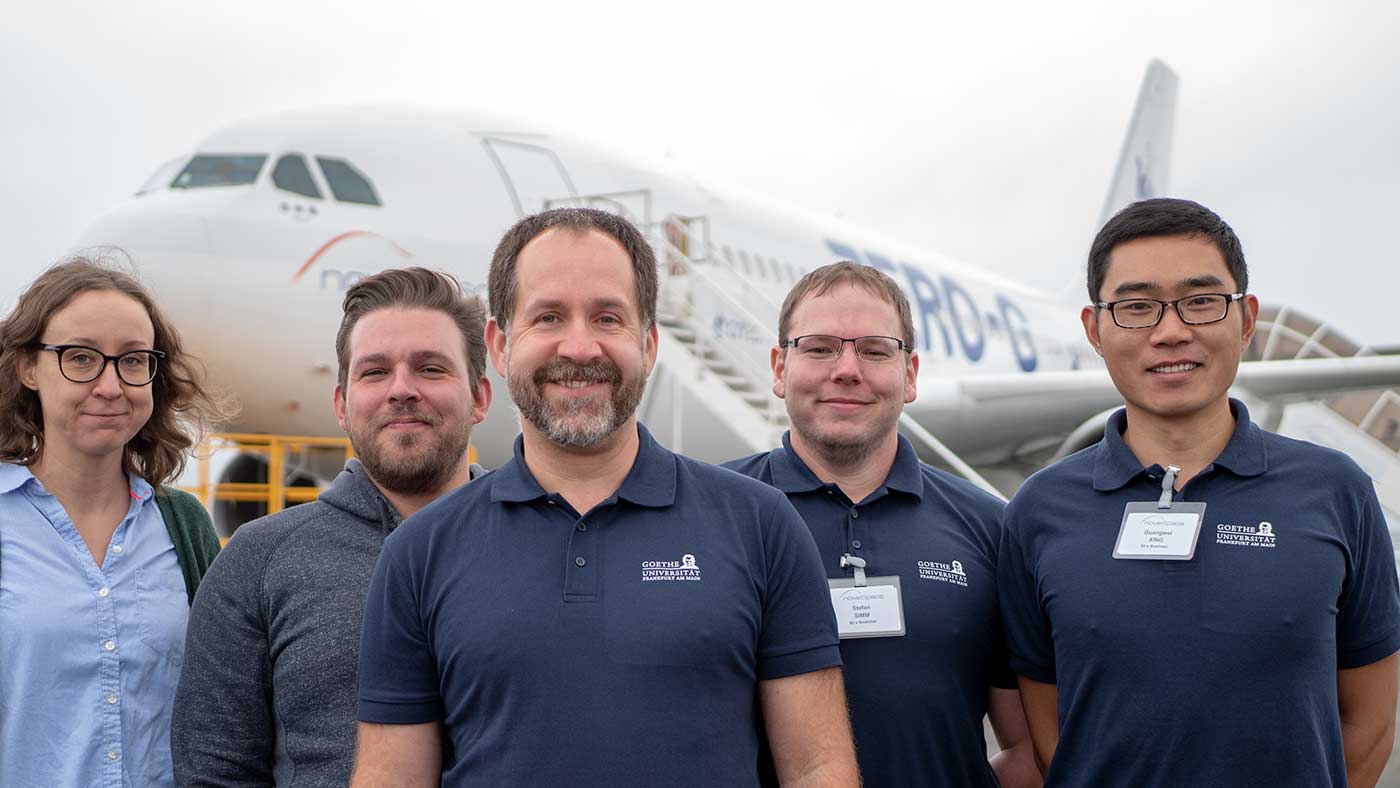
Zero-G lab

Due to their sessile lifestyle, an essential ability of plants is to adjust their growth towards or away from environmental stimuli. Plant responses that involve directed movements are called tropisms. Among the best-known tropisms are phototropism, the response to light, and gravitropism, the response to gravity. Gravity is one of the major factors that govern root growth in plants. Since the emergence of land plants, gravitropism allowed plants to adjust root growth to maximize access to water and nutrients and shoots to explore and exploit space on and above the surface of the Earth.
A particular focus of the group is on the nature of the gravireceptor, the role of early signaling pathways and secondary messengers, hormones, and the cytoskeleton. We perform experiments in altered gravity with the use of clinostats, centrifuges, and experimentation in the almost stimulus-free environment of microgravity provided by drop towers, parabolic flights of aircraft, sounding rockets, and low earth orbit space stations, like the International Space Station, which are increasingly contributing to our understanding of plant gravity sensing and orientation.
- Presse & Kommunikation
- Pressemitteilungen
- Öffentliche Veranstaltungen
- Uni-Publikationen
- Aktuelles Jahrbuch
- UniReport
- Forschung Frankfurt
- Karriere & Jobs
- Frankfurter Kinder-Uni
- Zahlen und Fakten
- Internationales
- Inbound: Aus der Welt nach Frankfurt
- Outbound: Von Frankfurt in die Welt
- Erasmus / LLP
- Goethe Welcome Centre (GWC)
- Refugees / Geflüchtete
- Erasmus +
- Sprachenzentrum oder Fremdsprachen
- Goethe Research Academy for Early Career Researchers
- Forschung
- Research Support
- Wissenschaftliche Zentren
- Unser Forschungsprofil
- Forschungsförderung
- Infrastrukturzentren
- Promovierende & Postdocs




