- GU Home
- Biowissenschaften
- Institute und Einrichtungen
- Institut MBW
- Arbeitskreise
- Abt. Müller-McNicoll
AK RNA Regulation in Higher Eukaryotes

Prof. Michaela Müller-McNicoll
Professor, Principal Investigator
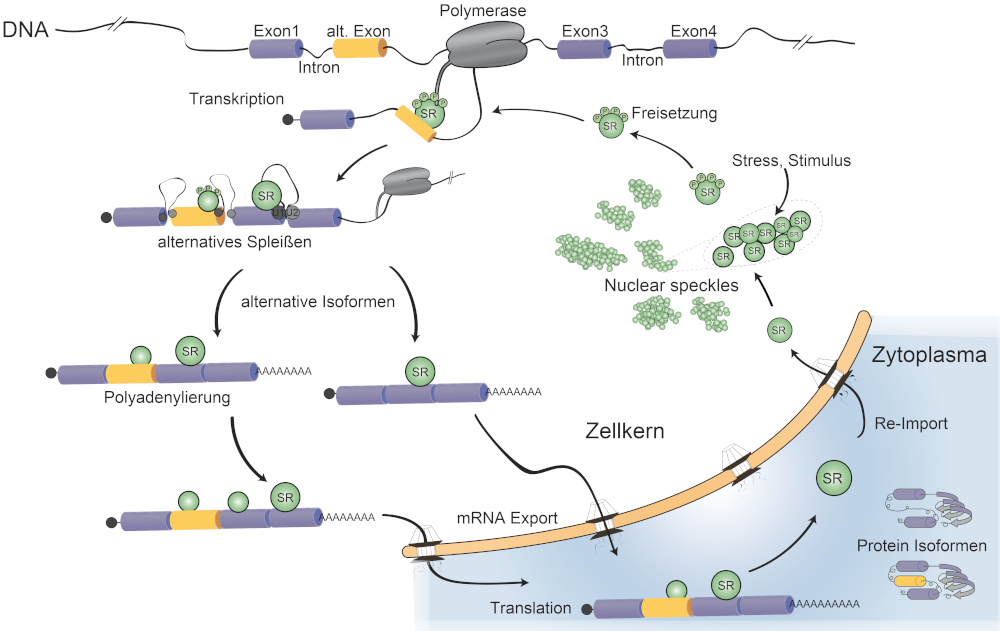
Eukaryotic mRNAs recruit numerous RNA-binding proteins (RBPs) and form large mRNA ribonucleoprotein particles called mRNPs. RBPs influence the fate of the mRNA at every step throughout its life cycle, from transcription of precursor mRNAs (pre-mRNAs) in the nucleus to translation and decay of mature mRNAs in the cytoplasm. Our group studies a family of nuclear RBPs, called SR proteins, which bind to specific sites in pre-mRNAs during transcription and regulate constitutive and alternative splicing (AS).
SR proteins bind mostly to coding exonic regions and are not removed during splicing. Therefore they have the potential to accompany mRNAs during their entire journey and indeed, some members of the SR protein family were shown to perform additional post-splicing functions in nuclear and cytoplasmic gene expression, regulating processes such as mRNA stability, microRNA processing, mRNA export or translation. However, the extent to which individual SR proteins are involved in post-splicing functions, whether these functions are connected to splicing and how such multifunctional and highly related RBPs can achieve specificity are not understood.
We aim to investigate whether and how SR proteins perform functions in post-splicing steps of gene expression, such as alternative polyadenylation (APA), nuclear surveillance, mRNA export and translation in the cytoplasm. Combining the use of different mammalian cells lines and BAC recombineering with diverse in vivo mRNP purification methods, genome-wide approaches and computational tools, we want to understand how sequence and structural information in the mRNA guides mRNP assembly and remodelling, how mRNP composition changes when the mRNA undergoes distinct maturation steps and how the quality of nuclear gene expression is secured in eukaryotic cells.
News
- January 2024: Ritaja and Harsh will join the lab as IMPRS PhD students. Welcome!
- December 2023: Our new paper is out in Journal of Cell Biology. Congratulations to Benni, Ricarda, Ellen, Frank and
- Irena. We developed and benchmarked a novel tool for the rapid degradation of GFP-tagged RNA binding proteins that localise to nuclear condensates.
- December 2023: Michaela was appointed as Max Planck Fellow at the MPI for Biophysics.
- November 2023: Johanna joins the lab as Postdoc. She will work in a joint project between the Schlundt group and our lab. Welcome!
- October 2023: Vanessa joins the lab as Master student. Welcome!
- October 2023: Vlado joins the group as Postdoc. He will wok in a joint project with the Schulz group and our lab. Welcome!
- September 2023: Benni successfully defended his PhD thesis. Congratulation
- May 2023: Tina joins the lab as Master student. Welcome!
- May 2023: The TRR267 ‘Noncoding RNAs in the cardiovascular system’ gets funded. We will receive funding for another 4 years. Congrats to Clara and Kathi.
- January 2023: Our new paper is out in Nucleic Acids Research. Congratulations to Camila, Clara, Michal, Frank, Nicole and Benni. We found that Poison cassette exon splicing of SRSF6 regulates nuclear speckle dispersal in hypoxia.
- August 2022: New collaborative paper with the group of Stefanie Dimmeler published on the role of the long non-coding RNA CALA in RNA decay.
- July 2022: Remus join the group as a Postdoc. He will work in a joint project between the Beck group and our lab. Welcome!
Kontakt
Arbeitskreis RNA Regulation in Higher Eukaryotes
Institut für Molekulare Biowissenschaften
Prof. Michaela Müller-McNicoll, PhD
Biologicum, Campus Riedberg
Gebäudeteil B, 1.OG
Max-von-Laue-Straße 13
60438 Frankfurt am Main
T +49 69 798 - 42079
Mueller-McNicoll(at)bio.uni-frankfurt.de
Sprechzeiten:
Donnerstags, 13-14 Uhr 13-14 Uhr nur nach Vereinbarung. Mindestens eine Woche im Voraus anSekretariat:
Frau Baechle Jourdan
office-muellermcnicoll@bio.uni-frankfurt.de
- Aktuelles und Presse
- Pressemitteilungen
- Öffentliche Veranstaltungen
- Uni-Publikationen
- Aktuelles Jahrbuch
- UniReport
- Forschung Frankfurt
- Aktuelle Stellenangebote
- Frankfurter Kinder-Uni
- Internationales
- Outgoings
- Erasmus / LLP
- Goethe Welcome Centre (GWC)
- Refugees / Geflüchtete
- Erasmus +
- Sprachenzentrum oder Fremdsprachen
- Goethe Research Academy for Early Career Researchers
- Forschung
- Research Support
- Forschungsprojekte, Kooperationen, Infrastruktur
- Profilbereich Molecular & Translational Medicine
- Profilbereich Structure & Dynamics of Life
- Profilbereich Space, Time & Matter
- Profilbereich Sustainability & Biodiversity
- Profilbereich Orders & Transformations
- Profilbereich Universality & Diversity