Molecular Evolution & Bioinformatics
Biosequence-Informatics
Prof. Dr. Ingo Ebersberger, ebersberger@bio.uni-frankfurt.de, Tel. +49 69 798 - 42112
Information
Biological sequences dominate the data basis for studies on organismic function and evolution. The improvement of high-throughput DNA sequencing methodologies meanwhile result in the availability of comprehensive genetic and genomic sequence information for organisms even from the remotest corner of the tree of life, and from species long extinct. Full exploitation of the information content of this data and correct interpretations are inextricably linked to four questions: How does one process, organise and analyse data sets from high-throughput DNA sequencing? What species are represented in the sequence data set? What can be learned from comparing present-day sequences about their evolutionary history and their function? And finally, what assumptions and (evolutionary) concepts underlie common bioinformatic sequence analysis algorithms, and how can these influence the outcome of an analysis?
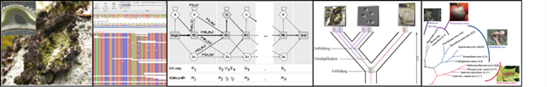
This module will use the assembly and analysis of a small eukaryotic genome as an example to address these questions. The practical work extends from the processing of raw sequence reads to the comparative analysis of the genes encoded in the reconstructed genome across a phylogenetically diverse set of representatives from the tree of life. Among others, we will investigate the evolutionary origins of the annotated genes, we will identify and interpret contaminating sequences in the genome assembly, and we will start predicting the likely functional consequences of gene loss. The practical exercises will be accompanied by lectures and a seminar in which the theoretical basics will be taught and deepened.
Start of the module- first half of the winter semester
Number of places available- 10
Special features - In consultation with the students, the module can be conducted entirely or partially in English.
- Presse & Kommunikation
- Pressemitteilungen
- Öffentliche Veranstaltungen
- Uni-Publikationen
- Aktuelles Jahrbuch
- UniReport
- Forschung Frankfurt
- Karriere & Jobs
- Frankfurter Kinder-Uni
- Zahlen und Fakten
- Internationales
- Inbound: Aus der Welt nach Frankfurt
- Outbound: Von Frankfurt in die Welt
- Erasmus / LLP
- Goethe Welcome Centre (GWC)
- Refugees / Geflüchtete
- Erasmus +
- Sprachenzentrum oder Fremdsprachen
- Goethe Research Academy for Early Career Researchers
- Forschung
- Research Support
- Wissenschaftliche Zentren
- Unser Forschungsprofil
- Forschungsförderung
- Infrastrukturzentren
- Promovierende & Postdocs




